In Your Genes: Single-cell RNA Sequencing can Help Understand and Treat Complex Diseases
Adverse reactions to drug treatment are an occasional, and often devastating, side effect of modern medicine. One particular condition called drug-induced hypersensitivity syndrome, or DiHS, is associated with treatment for bacterial and viral infections, and related diseases. Approximately 30% of all DiHS patients develop serious complications like infections and autoimmune disorders.
Generally, autoimmune disorders can be treated using synthetic corticosteroids (which decrease overall immune function), but some cases are resistant to treatment, posing a clinical challenge. Research on these cases is difficult, as DiHS cannot be studied using animal models, and with a mortality rate of roughly 8% it is often fatal, restricting studies in clinical settings as well.
 RNA, along with DNA, helps encode functional proteins, thus “guiding” overall cell function. Researchers can study RNA to uncover gene expression of viruses and other microbes, as well as in diseased human cells. (Photo courtesy: Shutterstock)
RNA, along with DNA, helps encode functional proteins, thus “guiding” overall cell function. Researchers can study RNA to uncover gene expression of viruses and other microbes, as well as in diseased human cells. (Photo courtesy: Shutterstock)
Professor Doyoung Kim MD PhD, an inflammatory disease specialist at the Yonsei University College of Medicine, South Korea, accidentally met a 44-year-old male patient suffering from DiHS. The patient had a severe skin rash with systemic symptoms that doctors couldn’t control. Any attempts to treat the rash resulted in severe flareups and progression to a “toxic epidermal necrolysis-like” presentation, worsening his condition. “I had experience working with cutting edge ‘single-cell RNA sequencing’ technology from my stint at Dr. Nagao’s laboratory (NIH, US) that studies basic skin immunology,” explains Prof Kim, “and with the challenging case of this DiHS patient, I saw an opportunity to utilize my knowledge in the hospital.”
RNA, like DNA, instructs cells how to “function,” in that it directs the synthesis of proteins that perform different functions in cells. Through the process of single-cell RNA, or scRNA, sequencing, researchers can figure out the pathophysiology of disease with great accuracy (see Manuscript figure below). This allows researchers to better study diseases without available animal models, such as DiHS.
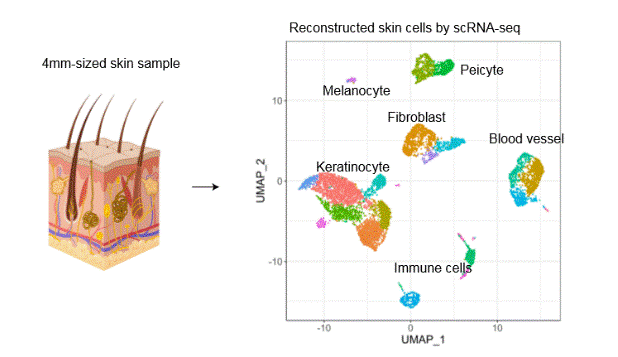 Manuscript figure: Using a tiny sample of skin tissue, scRNA-seq enables the measurement of gene expression in each individual cell. Each dot indicates an individual cell, and cells sharing similar biological characteristics lie adjacent on a UMAP.
Manuscript figure: Using a tiny sample of skin tissue, scRNA-seq enables the measurement of gene expression in each individual cell. Each dot indicates an individual cell, and cells sharing similar biological characteristics lie adjacent on a UMAP.
The research team led by Prof Kim “sequenced” the RNA of their DiHS patient’s skin and blood at a single cell level, identifying potential targets for drug application, as well as specific white blood cells that “held on” to the DNA information of the herpesvirus. Prof Kim states, “We initially aimed to find scientific clues for novel, personalized therapeutics for this patient. Interestingly, this new scRNA-seq technology itself indicated a new targetable cellular signaling pathway called ‘JAK-STAT’.”
With this new information, Professor Kim and his team were able to suppress the autoimmune response by targeting the JAK-STAT cellular signaling pathway and thus improve the patient’s symptoms caused by the adverse reaction. Prof Kim says, “Our study is the first to use scRNA-seq, a basic research technique, in real-world clinical practice. It has also successfully helped us find a new remedy for this rare, severe condition.”
The key discovery in this research is more than the application of one new therapy. “These successful results of scRNA sequencing will enable doctors to utilize it as a powerful tool for directing patient care in diseases with complex pathophysiology, including severe, rare inflammatory diseases and cancer, thereby making ‘personalized medicine’ possible,” concludes Prof Kim. This novel technique will improve the ability of medical staff to diagnose and treat challenging diseases, thereby improving patient recovery rate and quality of life.
Updated in July 2020
Recommended Articles
Professor Hyuk-Jae Chang
Professor Sung-Joo Hwang