A QUIRKY twist of fate: understanding epidermis cell differentiation in plants
To become a fully functioning multicellular organism much like ourselves, developing cells need to “know” what their jobs are, what they should look like, and where they should be located—a process called cellular differentiation. To understand how cells differentiate, scientists have used model organisms such as the plant Arabidopsis. In Arabidopsis, roots develop properly only after two kinds of epidermal cells (hair and non-hair) form in the appropriate locations. A group of researchers from Yonsei University and the University of Michigan has now updated our current understanding of how plant cell fate is determined.
In a study published in the journal Nature Communications, the scientists led by Dr. Myeong Min Lee “mutated” or manipulated Arabidopsis genes to understand the molecular signaling pathway involved in differentiation of root cells. Previously, scientists had figured out that non-hair cells form due to the expression of a gene called GL2. The protein WEREWOLF directly triggers GL2 expression, whereas the protein CAPRICE suppresses this expression. WEREWOLF and CAPRICE have an interesting interaction: in cells with WEREWOLF, the protein increases CAPRICE expression. CAPRICE then moves to adjacent cells to suppress WEREWOLF and GL2 expression. On top of that, the signaling molecule SCRAMBLED is required for this entire process and also seems to repress WEREWOLF in cells with CAPRICE. What we do not know is how SCRAMBLED is regulated. To understand that part of the pathway, Dr. Lee and colleagues decided to look at QUIRKY, a protein that interacts with SCRAMBLED for unclear reasons.
The researchers identified a mutant Arabidopsis plant harboring a mutation in the QUIRKY gene. The root cells in these plants were not properly developed, and experiments showed that WEREWOLF and CAPRICE genes were expressed in some epidermal cells regardless of their position, instead of being expressed in a position-dependent manner. Further experiments using plants with mutant QUIRKY and SCRAMBLED showed exactly how the two proteins interacted.
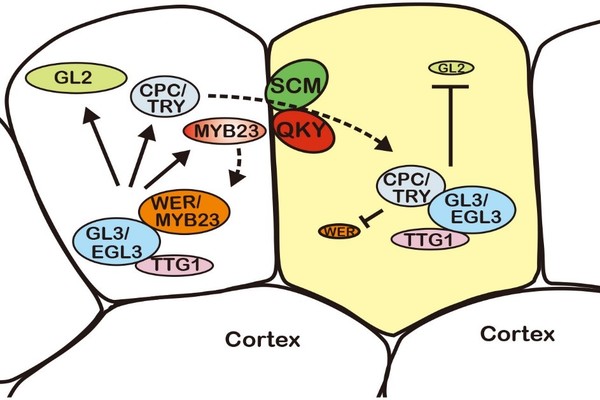 The protein QUIRKY (QKY) directly regulates the level of the protein SCRAMBLED (SCM) by preventing its degradation, which triggers the accumulation of another protein, called CAPRICE (CPC), in plant cells. This in turn suppresses the expression of the protein WEREWOLF (WER) and eventually leads to the formation of “hair cells,” which form the plant epidermis.
The protein QUIRKY (QKY) directly regulates the level of the protein SCRAMBLED (SCM) by preventing its degradation, which triggers the accumulation of another protein, called CAPRICE (CPC), in plant cells. This in turn suppresses the expression of the protein WEREWOLF (WER) and eventually leads to the formation of “hair cells,” which form the plant epidermis.
With regard to the results, Dr. Lee commented, “Basically, we showed that QUIRKY and SCRAMBLED work together to influence CAPRICE movement into the cells that become hair cells. Mechanistically, QUIRKY prevents addition of a protein called ubiquitin to SCRAMBLED and thus stops the latter’s breakdown by plant cells. This breakdown keeps SCRAMBLED from building up in certain cells, which allows WEREWOLF to work and trigger non-hair cell formation. So, when QUIRKY stops SCRAMBLED from degradation, it allows hair cells to form.”
To summarize, this study is the first to experimentally prove the role of QUIRKY in determining plant cell fate. “Our work shows how complex differentiation processes are even when only two cell types are involved,” Dr. Lee said. “On a theoretical level, we think these types of studies help paint a better picture of developmental processes in multicellular organisms. More practically, this knowledge can really help advance applied plant-science research, for instance, in the genetic engineering of crops.”
Updated in August 2019
Recommended Articles
Professor Jinwoo Cheon
m-Torquer: Magnetic Neuromodulation Could Make Us Real-Life ‘Magnetos’
Professor Jihyun F. Kim
Microbial Mercenaries for Plant Disease Resistance Ungrounded